- E-mail:BD@ebraincase.com
- Tel:+8618971215294
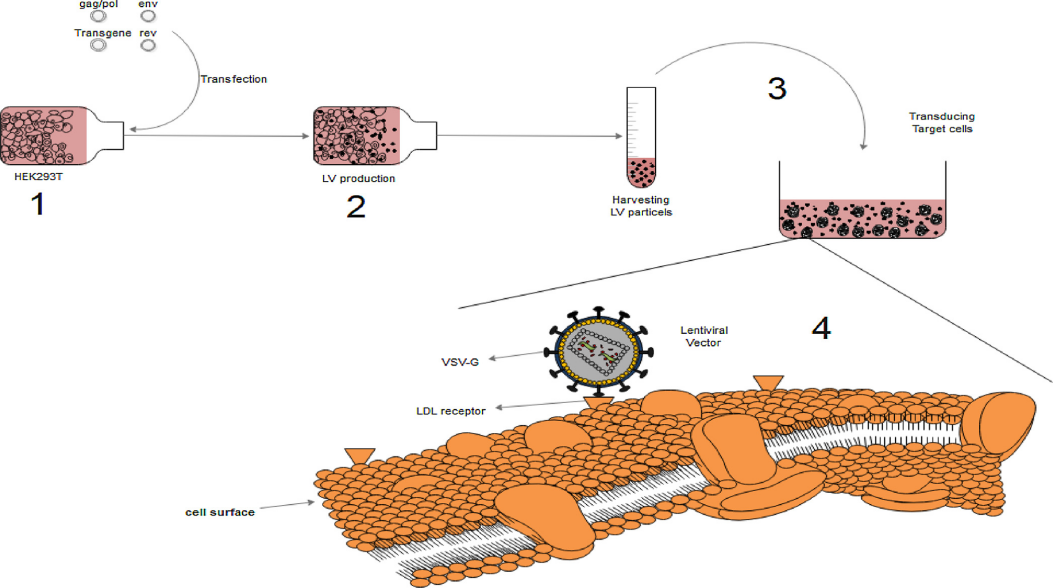
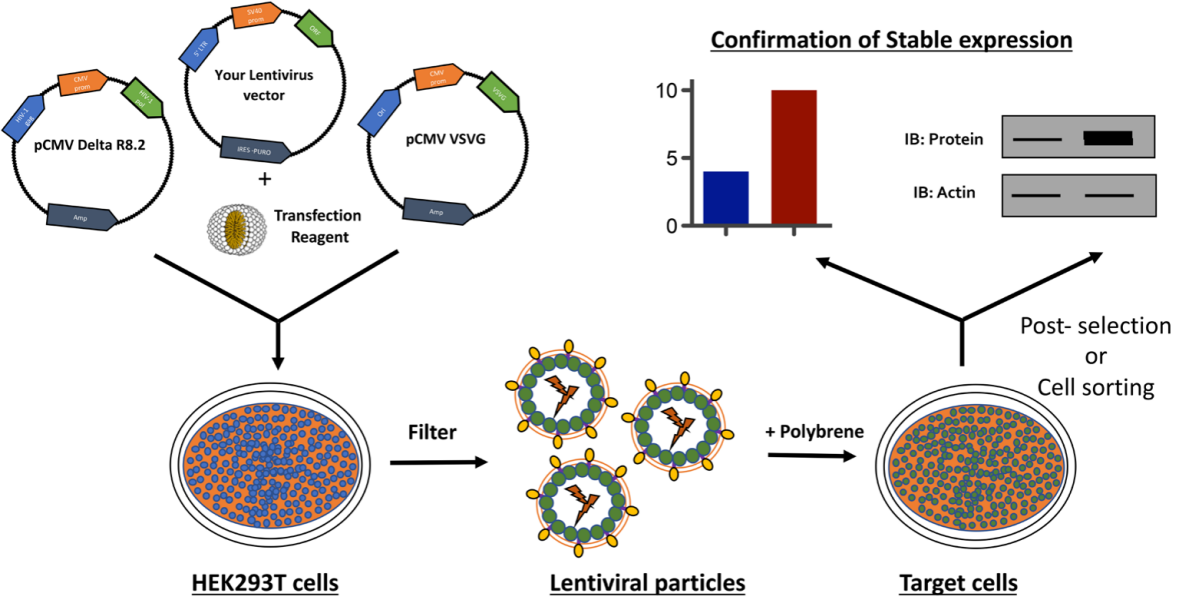
🔹Target Cell Preparation:
Select target cells in the logarithmic growth phase (optimal condition, highest transduction efficiency) and seed them into suitable culture containers (6-well plates/12-well plates, as required). Control the seeding density to ensure the cells reach 40%-60% confluency at the time of infection (too high a density may cause competition for nutrients, while too low a density may affect infection efficiency). Culture cells overnight in antibiotic-free complete medium and observe their condition the next day (only proceed if there is no contamination and cells are highly viable).
| Common Problem | Possible Causes | Solutions |
|---|---|---|
| Low Fluorescence Positive Rate (<30%) | 1. MOI too low; 2. Poor cell condition; 3. Insufficient virus activity; 4. Polybrene not added |
1. Increase MOI (do not exceed 100, with pre-experiment for toxicity); 2. Replace with target cells in logarithmic growth phase; 3. Use freshly thawed virus, concentrate if necessary; 4. Add 6-8 μg/mL Polybrene or use centrifugation method. |
| Massive cell death following infection (high toxicity) | 1. MOI too high; 2. Polybrene concentration too high; 3. Residual virus toxicity; 4. Cells are sensitive to the virus. |
1. Lower MOI, perform two rounds of infection (24 hours apart); 2. Reduce Polybrene concentration or omit it; 3. Replace the medium within 12 hours after infection; 4. Conduct a pre-experiment for virus toxicity to select the tolerated concentration. |
| No Surviving Cells After Selection or Slow Cell Growth | 1. Selection drug concentration too high; 2. Low infection efficiency, cells did not integrate the target gene; 3. Insufficient nutrients in the medium. |
1. Repeat the drug toxicity pre-experiment, reduce the selection drug concentration (set to 80% of MTC); 2. Optimize infection conditions to improve the positive rate, extend the pre-selection culture time; 3. Replace with fresh complete medium, add serum or growth factors. |
| Unstable Expression of Stable Cell Lines, Weak Fluorescence After Passage | 1. Incomplete selection, presence of mixed cells; 2. Poor integration site for the target gene; 3. Over-passage, cell aging. |
1. Use limiting dilution for clonal culture, purify the cell line; 2. Change lentivirus vector, optimize target gene sequence; 3. Passage stable cell lines no more than 20 generations, store in liquid nitrogen in time. |
| Bacterial/Fungal Contamination During Infection | 1. Non-sterile operation; 2. Virus solution or medium contamination; 3. Cells carrying bacteria. |
1. Terminate infection, thoroughly disinfect the experiment environment and equipment; 2. Replace with fresh culture medium and virus solution, trace the contamination source; 3. Discard contaminated cells, re-establish the experiment with bacteria-free cells. |
| No Difference in WB/qPCR Quantitative Detection (Overexpression/Knockdown Failure) | 1. Low infection efficiency, insufficient positive cell ratio; 2. Vector defects (gene cloning errors, incompatible promoters); 3. Improper timing of detection (expression/knockdown peak not reached); 4. Ineffective knockdown target or compensatory mechanisms; 5. Defective detection methods (poor primers/primary antibody, sample degradation). |
1. Verify fluorescence positive rate, re-optimize infection conditions if <50%; 2. Verify vector validity, change to compatible promoter or redesign shRNA target; 3. For overexpression, test qPCR 48-72 hours, WB 72-96 hours; for knockdown, test at 72-96 hours; 4. Design 3-4 shRNA targets to avoid compensation; 5. Optimize primers/primary antibodies, ensure no sample degradation, include positive control. |